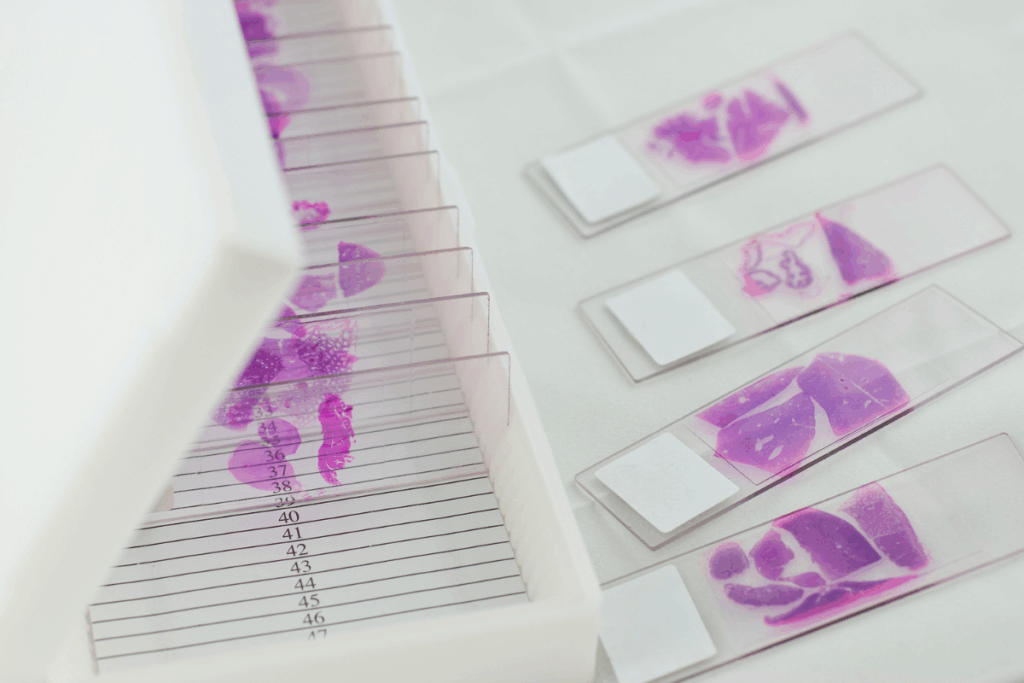
Advances in computational pathology and deep learning enable the analysis of histopathology images to predict tumor molecular characteristics. Models trained on H&E-stained whole-slide images aim to infer biomarkers relevant for cancer diagnosis and treatment, potentially reducing the need for expensive and time-consuming molecular testing. Studies indicate that these models can predict gene mutations and receptor status. They often exhibit inconsistent performance and limited generalizability across datasets because of the complex interplay of many molecular factors that affect tumor morphology.
The aim of this study was to assess the limitations of current deep learning methods used to predict molecular biomarkers from H&E-stained whole-slide images. The researchers sought to assess how interdependency among biomarkers and clinicopathological factors influences model prediction and to determine whether ignoring these relationships leads to biased or misleading performance. It also aimed to propose evaluation methods that can better detect biases and improve the reliability of machine learning models for precision oncology.
To address these objectives, the study analyzed biomarker relationships and machine learning model performance by using multiple large cancer datasets, including The Cancer Genome Atlas (TCGA), METABRIC, Memorial Sloan Kettering (MSK), and Dana-Farber Cancer Institute (DFCI) cohorts. These datasets contain paired histopathology images and molecular information, which allow researchers to train and evaluate deep learning models for biomarker prediction. The researchers examined molecular interactions in biomarkers by assessing patterns of mutual exclusivity and co-occurrence, which are common in cancer biology. For example, certain mutations frequently occur together, while others rarely appear in the same tumor.
The study evaluated how these relationships influence model predictions by using statistical techniques like permutation testing and stratification analysis. Models were tested across subgroups defined by the presence or absence of other biomarkers to determine whether predictive performance remained stable. The researchers examined the influence of clinicopathological variables, which include tumor grade and tumor mutational burden (TMB), which are known to affect tumor morphology and may act as confounding variables. By analyzing model performance within grade- and TMB-stratified subgroups, the study assessed whether models relied on these features as proxies instead of learning the intended biomarker signals.
The analysis revealed significant interdependencies among biomarkers in different cancer types and datasets. These relationships appeared as patterns of mutual exclusivity or co-occurrence and reflected both biological mechanisms and cohort-specific associations. The study found that many deep learning models failed to account for these dependencies during training. Consequently, model predictions for a specific biomarker were often influenced by the status of other related biomarkers. For instance, the performance of a progesterone receptor (PR) prediction model dropped dramatically in tumors with CDH1 mutations, with the area under the receiver operating characteristic curve (AUROC) decreasing from 0.79 to 0.50. This finding indicated that the model could not distinguish the independent signal of the PR biomarker when another related mutation was present. Similar issues were observed in colorectal cancer when predicting MSI and BRAF mutations. Because MSI-high tumors frequently harbor BRAF mutations, models often confuse the two biomarkers.
Although models sometimes achieved high overall accuracy, their performance declined when evaluated in specific subgroups defined by other biomarkers. The study also showed that models frequently relied on morphological features linked to tumor grade or tumor mutational burden instead of true biomarker signals. Predictions for estrogen receptor (ER) status appeared highly accurate in external datasets because of a stronger association between tumor grade and ER status, rather than the model learned a genuine genotype-phenotype relationship. When evaluated within grade-stratified groups, model performance was comparable to that of simple grade-based classifiers. These findings indicate that current models may produce misleading results when applied to different patient populations or clinical contexts.
This study highlights limitations in current deep learning models that predict molecular biomarkers from histopathology images, noting that ignoring intricate molecular and clinical interactions can result in biased predictions and unreliable outcomes. These models are not ready to replace standard genomic testing in clinical practice; they can serve as valuable tools for screening and research when used along with confirmatory testing. Further research on causal machine learning methods is needed to improve accuracy and reduce confounding. Improved methodologies and diverse datasets can lead to better precision diagnostics in oncology.
Reference: Dawood M, Branson K, Tejpar S, et al. Confounding factors and biases abound when predicting molecular biomarkers from histological images. Nature Biomedical Engineering. Published March 2, 2026. doi:10.1038/s41551-026-01616-8